To understand the different sequence alignment algorithms that are present in MacVector, please check out this blog post. For more information on the use of the Align To Reference function, please check out the Align To Reference – Sequence Confirmation Tutorial.pdf tutorial that you will find installed in the Applications/MacVector/Documentation/ folder. Note that the algorithm automatically “flips” reads to maintain the best alignment and will “clip” out vector sequences (shown in gray text in the image). Click on the Align button, choose the Sequence Confirmation algorithm and click OK to align all of the reads against the reference.You can also use plain sequence files in any format supported by MacVector. Click on the Add Seqs button and select the ABI (.ab1) files sent by your sequencing facility.This might be the PCR fragment you have cloned, or the construct you are re-sequencing. However, this suffers from a couple of limitations: (a) ClustalW does not know how to automatically “flip” sequences when you have pairs of reads on opposite strands and (b) the MSA interface doesn’t let you view the raw trace data to help identify sequencing artifacts.Ī far better solution is to use the Analyze | Align To Reference function which is tuned for exactly this type of analysis Many users immediately think “this is a multiple sequence alignment problem” and so they attempt to align the sequences sent back from the sequencing facility using ClustalW and the Multiple Sequence Alignment interface. Perhaps it is a cloned PCR fragment where you want to confirm the sequence, perhaps you are sequencing across a cloning junction, or maybe screening clones for a successful mutagenesis experiment. 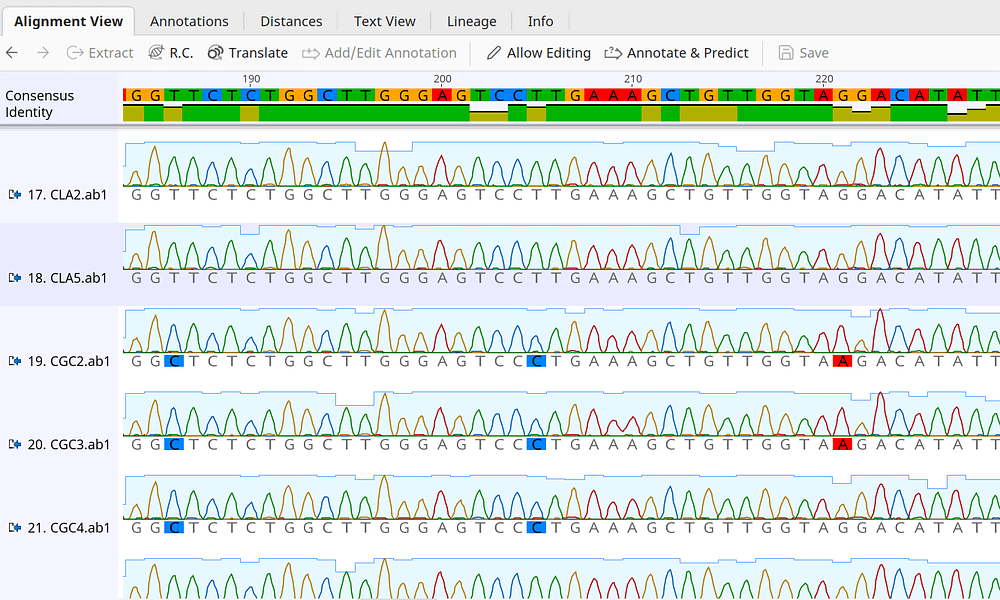
You can learn more about sequence alignments on the UniProt help page.One of the most common tasks in any molecular biology lab is the need to re-sequence a piece of DNA. You can also run Alignment from within the Basket. All relevant results pages (such as UniProtKB, UniRef, UniParc and tool results) provide an ‘Align’ button to run alignments directly by selecting entries with checkboxes. The following kinds of UniProt identifiers are supported: P00750Įach UniProtKB entry which contains both a sequence and one or more isoforms of that sequence, enables you to align the canonical sequence and its isoforms. Note – advanced users are given the option of varying the alignment parameters from those given as default. Enter either protein sequences in FASTA format or UniProt identifiers (as above) into the form field.Click on the Align link in the header bar to align two or more protein sequences with the Clustal Omega program.Exercise: mapping other database identifiers to UniProtĪll materials are free cultural works licensed under a Creative CommonsĪttribution 4.0 International (CC BY 4.0) license, except where further licensing details are provided.Ī sequence alignment is a way of arranging the primary sequences of a protein to identify regions of similarity that may be a consequence of functional, structural, or evolutionary relationships between the sequences.Exercise: finding entries with 3D structures.Downloading a proteome set for specific organism.Accessing UniProt data programmatically. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
